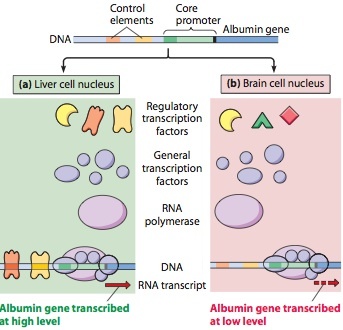
What are the 5 main levels in which gene control is exerted?
- the genome,
- transcription,
- *RNA processing and export from nucleus to cytoplasm,
- *translation,
- *posttranslational events.
*Regulatory mechanisms in the last three categories are all examples of posttranscriptional control
The haploid genome of Xenopus normally contains about 500 copies of the genes that code for 5.8S, 18S, and 28S rRNA. However, the DNA of these genes is selectively replicated about 4000-fold during oogenesis (development of the egg prior to fertilization). (B:723)
Why?
the mature oocyte contains about 2 million copies of the genes for rRNA. This level of amplification is apparently needed to sustain the enormous production of ribosomes that occurs during oogenesis, which in turn is required to sustain the high rate of protein synthesis needed for early embryonic development.
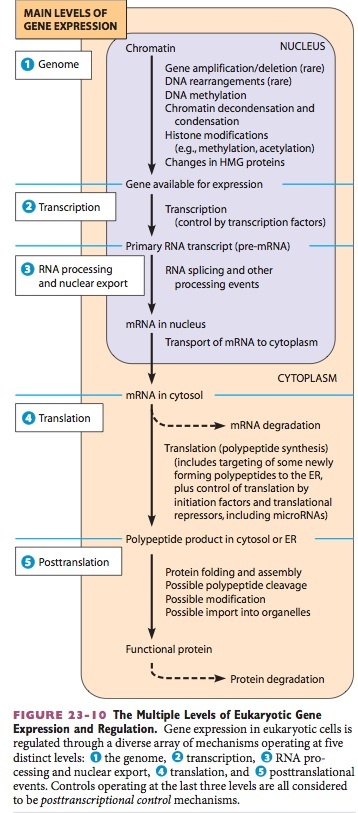
Sensitivity of Active Genes in Chromatin to Digestion with DNase I
DNase I hypersensitive sites
- When nuclei are treated with very low con- centrations of DNase I, it is possible to detect specific locations in the chromatin that are exceedingly susceptible to digestion.
- tend to occur up to a few hundred bases upstream from the tran- scriptional start sites of active genes
- about ten times more sensitive to DNase I digestion
- correspond to regions in which the DNA is not part of a nucleosome.
DNA methylation
- addition of methyl groups to selected cytosine bases in DNA.
- Methylation of promoter regions can either block access of proteins required for transcriptional acti- vation or serve as a binding site for proteins that condense chromatin into inactive configurations.
- The net effect is either a localized or regional silencing of gene expression.
- inheritable epigenetic changes
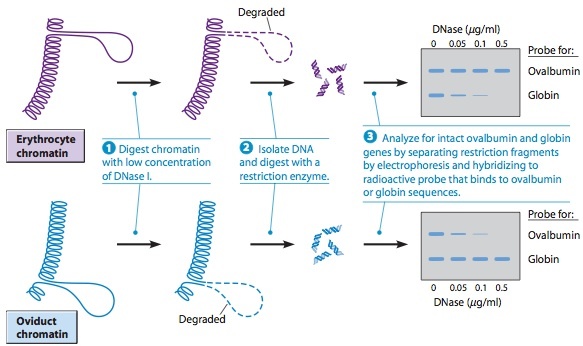
Roles of Histone Acetylation and Chromatin Remodeling Proteins during Transcriptional Activation.
Repressor proteins, which cause transcription to occur less frequently at a particular site, recruit HDAC complexes, and activator proteins recruit HAT complexes.
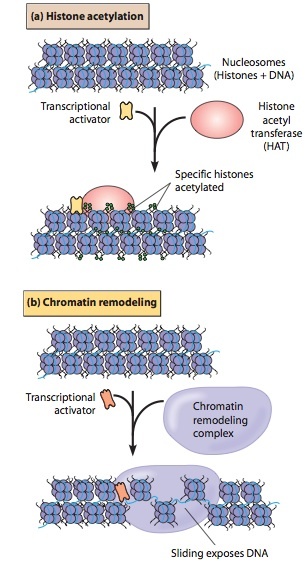
(a) Transcriptional activator proteins recruit histone acetyltrans- ferase (HAT) complexes to the region near a gene. These enzymes then add acetyl groups, which leads to the opening or closing of the chromatin structure in the region.
(b) Chromatin remodeling proteins can have several effects on nucleosomes. In the case shown here, a chromatin remodeling protein causes sliding of nucleosomes, exposing a region of DNA that could then be transcribed.
nuclear run-on transcription
provides a snapshot of the transcriptional activity occurring in a nucleus at a given moment in time
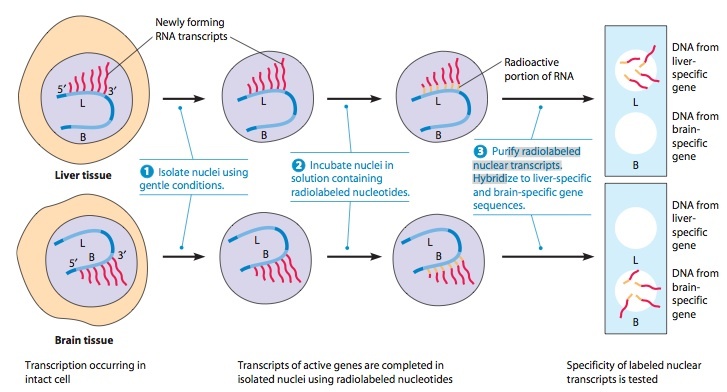
Demonstration of Differential Transcription by Nuclear Run-on Transcription
Assays
(B: Fig 28-18)
In these studies, B represents a hypothetical gene that is expressed only in brain tissue, and L is a gene expressed only in liver tissue. Isolated nuclei are incubated in a solution containing radioactively labeled ribonucleotides, which become incorporated into the mRNA being synthesized by active genes. If different genes are active in liver and brain tissue, some labeled sequences in the liver nuclear transcripts will not be present in brain transcripts and vice versa. The composition of the labeled RNA population is assayed by allowing the labeled RNA to hybridize with DNA sequences representing different genes that have been attached to a filter-paper support ( 3 ). Labeled liver transcripts hybridize with a different set of genes than do labeled brain transcripts, indicating that the identities of the active genes in the two tissues differ.
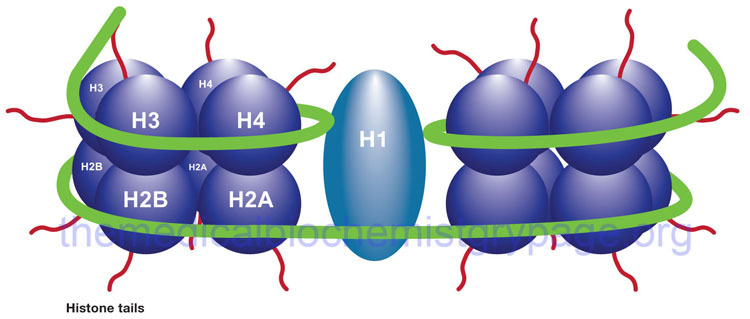
Diagrammatic representation of a nucleosome
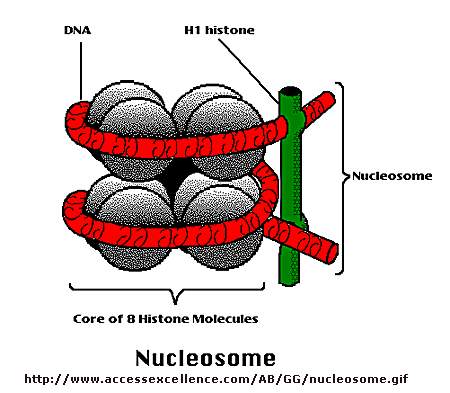
Diagrammatic representation of a nucleosome (smaller)
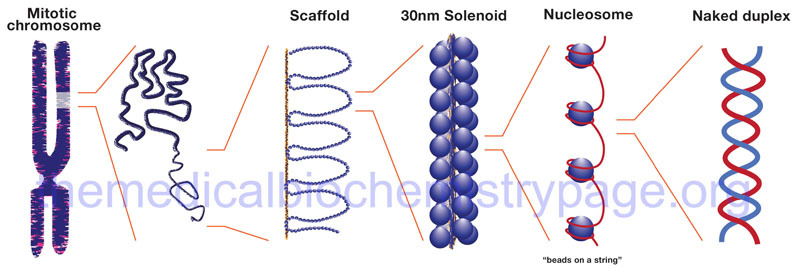
Hierarchy of chromatin structure
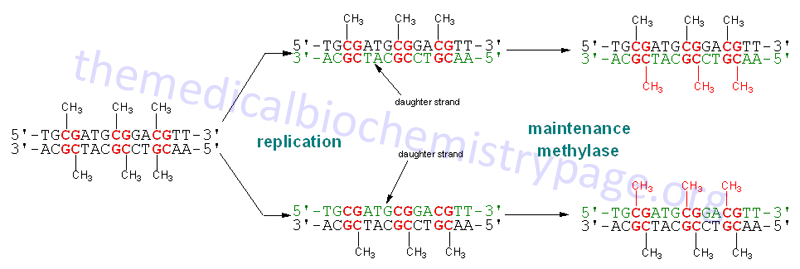
Post-Replicative Modification of DNA, Methylation
http://themedicalbiochemistrypage.org/dna.html#chromatin
proximal control elements
- stimulate transcription of the gene by interacting with regulatory transcription factors
- sequences located upstream of the core promoter but within about 100–200 base pairs of it.
- number, identity, and exact location vary gene to gene
regulatory transcription factors.
- Transcription factors that selectively bind to one of the proximal control elements, or to other control sequences located outside the core pro- moter,
- increase (or sometimes decrease) transcription initiation by interacting with components of the transcription apparatus.
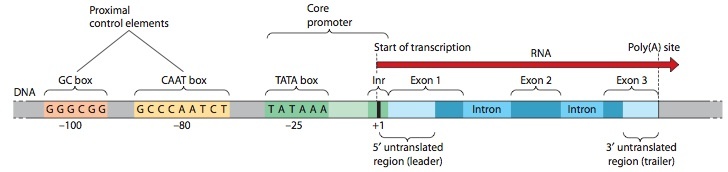
Anatomy of a Typical Eukaryotic Gene, with Its Core Promoter and Proximal Control Region
This diagram (not to scale) features a typical protein-coding eukaryotic gene, which is transcribed by RNA polymerase II. The promoter—called the core promoter to distinguish it from the proximal control region—is characterized by an initiator (Inr) sequence surrounding the transcriptional startpoint and a TATA box located about 25 bp upstream (to the 5¿ side) of the startpoint. The core promoter is where the general transcription factors and RNA polymerase assemble for the initiation of transcription. Within about 100 nucleotides upstream from the core promoter lie several proximal control elements, which stimulate transcrip- tion of the gene by interacting with regulatory transcription factors. The number, identity, and exact location of the proximal elements vary from gene to gene. Here we show a simple case involving one copy of each of two common elements, the GC box and the CAAT box. The transcription unit includes a 5’ untranslated region (leader) and a 3’ untranslated region (trailer), which are transcribed and included in the mRNA but do not contribute sequence information for the protein product. At the end of the last exon is a site where, in the primary transcript, the RNA will be cleaved and given a poly(A) tail.
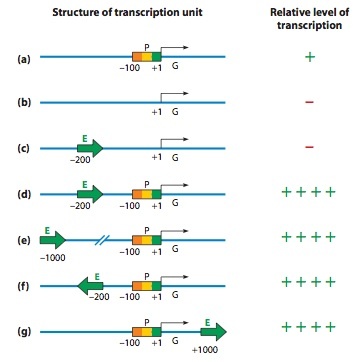
Effects of Enhancer Orientation and Location
Recombinant DNA techniques have been used to alter the orientation and location of DNA control elements and study the effect of such changes on gene transcription. The black arrows indi- cate the direction of transcription of gene G, with the startpoint (first transcribed nucleotide) labeled +1. The other numbers give the positions of nucleotides relative to the startpoint. (a) The core promoter (P) alone, in its typical location just upstream of gene G, allows a basal level of transcription to occur. (b) When the core promoter is removed from the gene, no transcription occurs.
(c) An enhancer (E) alone cannot substitute for the promoter region, but (d) combining an enhancer with a core promoter results in a significantly higher level of transcription than occurs with the promoter alone. (e) This increase in transcription is observed when the enhancer is moved farther upstream, (f) when it is inverted in orientation, and (g) even when it is moved to the 3’ side of the gene.
core promoter
- characterized by an initiator (Inr) sequence surrounding the transcriptional startpoint and a TATA box located about 25 bp upstream (to the 5’ side) of the startpoint
- where the general transcription factors and RNA polymerase assemble for the initiation of transcription
transcription unit
5’ untranslated region (leader) and a 3’ untranslated region (trailer), which are transcribed and included in the mRNA but do not contribute sequence information for the protein product.
silencers/repressors
Silencers share many of the features of enhancers, except that they inhibit rather than activate transcription. Because the binding of regulatory transcription factors to silencers reduces rather than increases gene transcription rates
insulators
Insulator sequences, along with their associated binding proteins, create physical barriers between neighboring DNA regions that prevent enhancers or silencers from exerting their effects across the barrier.
One complication encountered with silencers and enhancers stems from their ability to influence the tran- scription of faraway genes, which could be problematic if genes with opposing functions reside in neighboring regions. For example, a group of genes active in one cell type might lie near another set of genes that should not be active in those same cells. In such situations, DNA sequences called insulators are sometimes employed to prevent an enhancer (or silencer) from inadvertently acting on both groups of genes simultaneously.
Two basic principles govern the interaction between enhancers and the genes they regulate
1) looping of the DNA molecule can bring an enhancer into close proximity with a promoter, even though the two lie far apart in terms of linear dis- tance along the DNA double helix.
2) a diverse group of coactivator proteins mediate the interaction between activators bound to the enhancer and the RNA polymerase complex associated with the promoter.
Mediator
functions as a coactivator by serving as a “bridge” that binds to activator proteins associated with the enhancer and to RNA polymerase, thereby linking enhancers to the components involved in initiating transcription at RNA polymerase II promoters.
Mediator serves as a central coordinating unit for gene regulation, receiving both positive and negative inputs and transmitting the information to the transcription machinery.
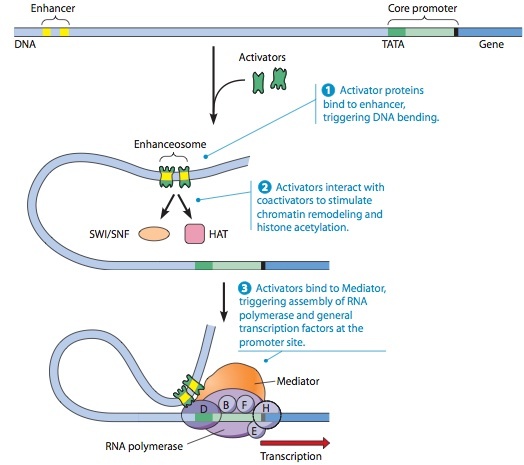
A Model for Enhancer Action.
In this model, an enhancer located at a great distance along the DNA from the protein-coding gene it regulates is brought close to the core promoter by a looping of the DNA.
- Regulatory transcription factors called activators first bind to the enhancer elements, triggering DNA bending that brings the activators closer to the core promoter.
- The activators then interact with coac- tivator proteins such as SWI/SNF, which causes chromatin remodeling, and HAT (histone acetyltransferase), which catalyzes histone acetylation. The net effect is to decondense the chromatin and make the DNA in the promoter region more accessible.
- The activators then bind to another coactivator called Mediator, and the activator-Mediator complex facilitates the correct positioning of general transcription factors and RNA poly- merase at the promoter site, allowing transcription to be initiated. For simplicity, the figure is drawn with only two activators, but often half a dozen or more are involved.
combinatorial model for gene regulation
- proposes that a relatively small number of different DNA control elements and transcription factors, acting in different combinations, can establish highly specific and precisely controlled patterns of gene expression in dif- ferent cell types.
- assumes that some transcription factors are present in many cell types. including general transcription factors, required for transcription in all cells, plus any regulatory factors needed for transcribing constitutive genes and other genes that are frequently expressed.
- In addition, transcription of genes that encode tissue-specific proteins requires the presence of transcription factors or combinations of tran-scription factors that are unique to individual cell types