recombinant DNA molecules
constructed from pieces of naturally occurring chromosomes and plasmids
Module 3
polymerase chain reaction (PCR)
- repeated copying of a short segment of a DNA molecule
- uses 2 primers
- 3’ end of primers are facing each other
The basis of recombinant DNA technology is the ability to manipulate DNA molecules in the test tube. This, in turn, depends on the availability of _____ ______ whose activities are known and can be controlled, and which can therefore be used to make specified changes to the DNA molecules
purified enzymes
The enzymes available to the molecular biologist fall into four broad categories:
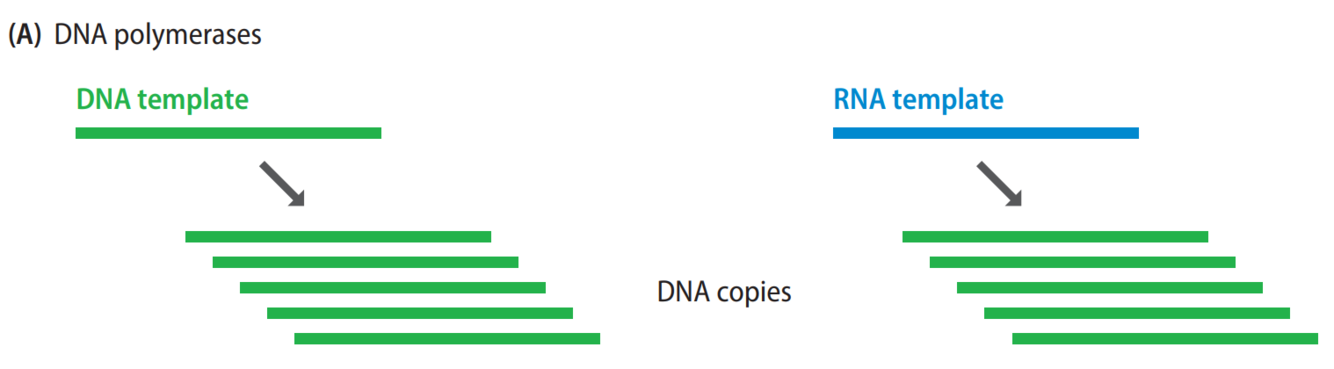
- DNA polymerases: synthesize new polynucleotides complementary to an existing DNA or RNA template
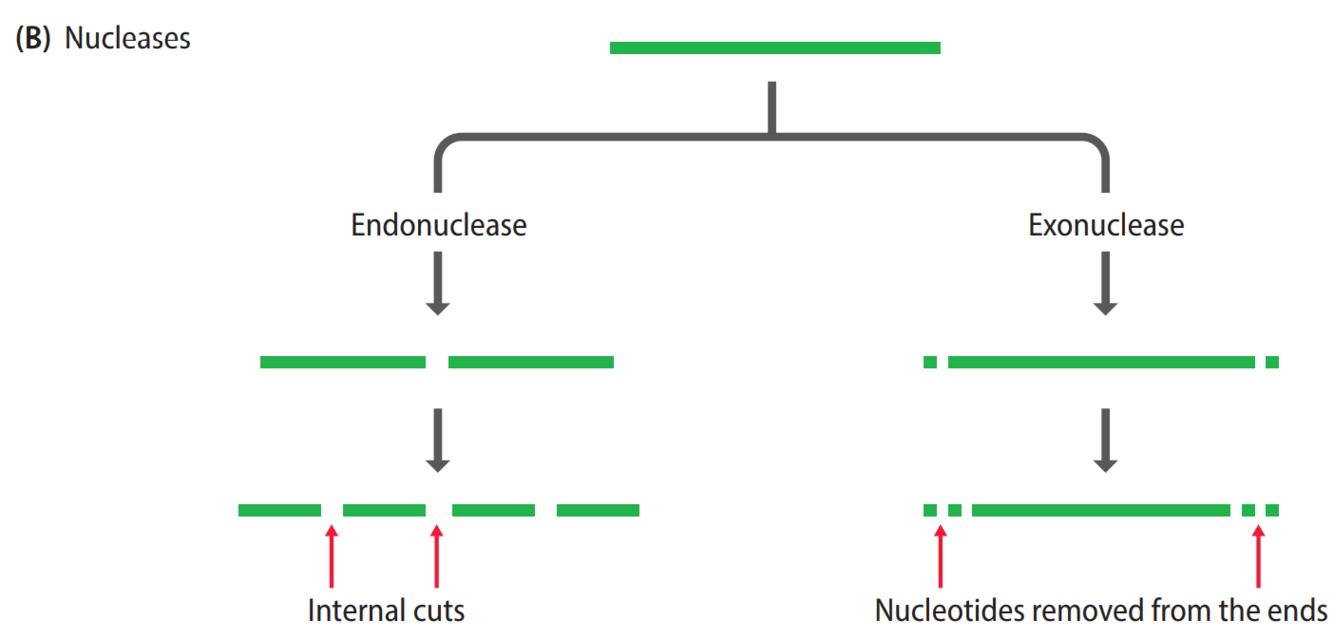
- Nucleases: degrade DNA molecules by breaking the phosphodiester bonds that link one nucleotide to the next
- Ligases: join DNA molecules together by synthesizing phosphodiester bonds between nucleotides at the ends of two different molecules or at the two ends of a single molecule
- End-modification enzymes: make changes to the ends of DNA molecules
Module 3
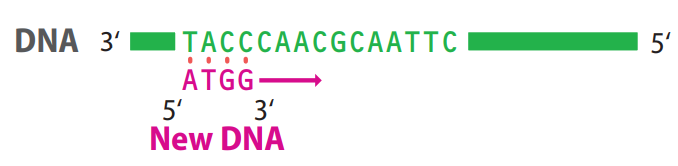
The new polynucleotide is always synthesized in the _____ direction: DNA polymerases that make DNA in the other direction are unknown in nature.
5ʹ → 3ʹ


Module 3
- DNA polymerase requires a _____ in order to initiate the synthesis of a new polynucleotide.
- The sequence of this primer determines the … and …
- When a DNA polymerase is used to make new DNA in vitro, the primer is usually a ______ _______ made by chemical synthesis.
- The primer is usually ______ nucleotides in length
- 3’ mismatch =
- Mismatches at other positions are _____ and useful for …
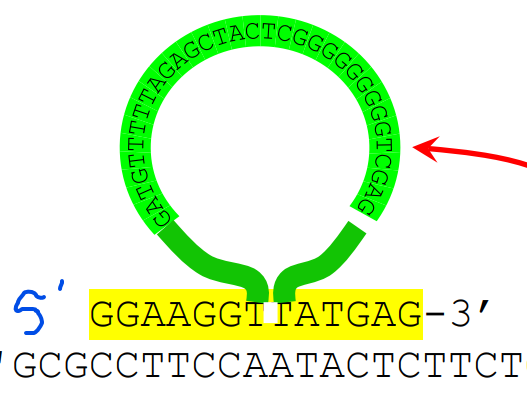
- ____ _____ within primer are tolerated, useful for introducing ______ into genes In the same way, deletions in primer are useful for introducing _____ into genes
- primer
- position at which it attaches to the template DNA
- specifies the region of the template that will be copied
- short oligonucleotide
- 20
- no PCR product, because DNA polymerase can’t extend
- tolerated
- introducing mutations into genes
- Large insertions
- insertions
- deletions
Module 3
the DnA synthesis and exonuclease activities of DnA polymerases
- 5ʹ → 3ʹ DNA synthesis activity: enables polymerase to add nucleotides to the 3’-end of the strand that it is synthesizing
- 3ʹ → 5ʹ exonuclease activity: enables polymerase to remove one or more nucleotides from the 3’-end of the strand that it is making (proofreading)
- 5ʹ → 3ʹ exonuclease activity: enables polymerase to remove one or more nucleotides from the 5ʹ-end of a polynucleotide that is already attached to the template strand.
- can degrade the 5ʹ-end of a polynucleotide that has just been synthesized
- enzyme is almost always prepared from E. coli cells whose polymerase gene has been engineered so that the resulting enzyme has the desired properties
- aka Gap Repair
- Usually during synthesis of lagging strand
- Cleavage of the phosphodiester bond of DNA
exonucleases
removes nucleotides from the ends of DNA and/or RNA molecules
endonucleases
makes cuts at internal phosphodiester bonds
Module 3
restriction endonucleases types
- three main types: I, II, III
- With types I and III, there is no strict control over the position of the cut
- with type II the cut is always at the same place, either within the recognition sequence or very close to it
restriction endonucleases
- enzyme that binds to a DNA molecule at a specific sequence
- makes a doublestranded cut at or near that sequence
- Because of sequence specificity, the positions of cuts within a DNA molecule can be predicted if the DNA sequence is known
There are also examples of restriction enzymes with degenerate recognition sequences, meaning that
- they cut DNA at any of a family of related sites
- i.e. HinfI recognizes 5ʹGANTC3ʹ, where N is any nucleotide, and so it cuts at 5ʹGAATC3ʹ, 5ʹGATTC3ʹ, 5ʹGAGTC3ʹ, and 5ʹGACTC3ʹ
- Most cut within the recognition sequence, but a few, such as BsrBI, cut at a specified position outside of this sequence
Module 2
Restriction enzymes cut DNA in two different ways:
- blunt or flush end: simple doublestranded cut
- sticky or cohesive ends:
- cuts the two DNA strands at different positions, usually two or four nucleotides apart
- not good for creating genomic libraries because larger overhangs of at least 15 to 16 bp are needed to ensure a single occurrence in human genome
- resulting DNA fragments have short, singlestranded overhangs at each end
- base pairing between them can stick the DNA molecule back together again
One feature that is particularly important in recombinant DNA technology is that some pairs of restriction enzymes have different _____ ______ but give the same _____ _____
- recognition sequences
- sticky ends
Module 3
gel electrophoresis
the standard method for separating DNA molecules of different lengths
in gel electrophoresis. _____ _____ is the critical determinant of migration rate. This is because …
- molecular length
- the gel is a network of pores through which the DNA molecules have to travel to reach the positive electrode
- Shorter molecules are less impeded by the pores than are longer molecules and so move through the gel more quickly
In order to follow the progress of the electrophoresis, one or two _____ of known migration rates are added to the DNA samples before loading. The bands of DNA can be visualized by soaking the gel in _____ ______ solution. This compound ______ between DNA base pairs and ______ when activated with _____ ______
- dyes
- ethidium bromide
- intercalates
- fluoresces
- ultraviolet radiation
DNA fragments that have been generated by treatment with a restriction endonuclease can be joined back together again or attached to a new partner by a DNA ligase. The reaction requires energy, which is provided by
adding either ATP or nicotinamide adenine dinucleotide (NAD) to the reaction mixture, depending on the type of ligase used
Module 3
quantitative PCR
- enables the amount of target DNA present at the start of the PCR to be measured
- rate of product synthesis during the test PCR is compared with the progress of control PCRs with known amounts of starting DNA
- comparison imade by identifying the stage in the PCR at which the amount of fluorescent signal reaches a preset threshold
- The more rapidly the threshold is reached, the greater the amount of target DNA in the starting mixture
Module 3
In a PCR experiment, the target DNA is mixed with
- Taq DNA polymerase
- pair of oligonucleotide primers
- nucleotides
Module 3
Primers attach to the target DNA at ____ _____ of the segment to be copied. This means that the sequences of these attachment sites must be known so that primers of the appropriate sequences can be _____
- either side
- synthesized
Module 3
Reaction is started by heating the mixture to ______ so that the
_____ ______ that hold together the two polynucleotides of the double helix are broken, _____ the target DNA into single-stranded molecules
- 94°C
- hydrogen bonds
- denaturing









