Module 5
Homologous genes
- share a common evolutionary ancestor, revealed by sequence similarities between the genes
Module 5
Orthologous genes
- homologs present in different organisms
- common ancestor predates the split between the species
- usually have the same or very similar functions
Module 5
Paralogous genes
- present in the same organism
- often as members of a recognized multigene family
- common ancestor may or may not predate the species in which the genes are now found
Module 5
A homology search can be conducted with a _____ sequence, but usually a tentative gene sequence is converted into an _____ _____ sequence before the search is carried out
- DNA
- amino acid
Module 5
Why in a homology search, is the sequence converted into an amino sequence?
- there are 20 different amino acids in proteins
- only four nucleotides in DNA
- genes that are unrelated usually appear to be more different from one another when their amino acid sequences are compared
Module 5
how is a homology search performed
- by making alignments between the query sequence and sequences from the databases
- For each alignment, a score is calculated
- two ways of generating the score
- simplest programs
- count the number of positions at which the same amino acid is present in both sequences
- when converted into a percentage, gives the degree of identity between two sequences
- uses chemical relatedness between nonidentical amino acids to assign a score
- higher score for identical or closely related amino acids
- lower score for less closely related amino acids
- To achieve the highest possible score, the algorithm introduces gaps at various positions in one or both sequences
- parallels processes thought to occur during the evolution of genes
Module 5
standard BLAST program is efficient at identifying homologous genes that have more than _____% sequence similarity but is less effective at recognizing evolutionary relationships if the similarity is _____
- 40
- lower
Module 5
PSI-BLAST (position-specific iterated BLAST)
identifies more distantly related sequences by combining the homologous sequences from a standard BLAST search into a profile, the features of which are used to identify additional homologous sequences that were not detected in the initial search.
Problems with BLAST
the presence in the databases of genes whose stated functions are incorrect.
may be possible to deduce at least some part of the function of the gene by searching the amino acid sequence for _____ that encode _____ _____ of known function
- motifs
- protein domains
Zinc fingers are _____-_____ structures, so identification in an unknown gene of an amino acid sequence that can encode a zinc finger indicates that the gene codes for
- DNA-binding
- a DNA-binding protein
searching for motifs called _____ _____, which direct proteins to organelles such as the nucleus or mitochondria or might specify that the protein is secreted from the cell
sorting sequences
The presence of a shared domain indicates that two proteins can perform a similar _____ _____, but that does not necessarily mean that the proteins have
- biochemical activity
- similar overall functions
Identification of a domain sequence in an unknown gene therefore identifies a specific ____ _____ to be identified, but on its own this does not enable the actual _____ of the gene to be assigned
- biochemical activity
- function
Module 5
If the starting point is the gene, rather than the phenotype, then one strategy is
- to mutate the gene and identify the phenotypic change that results
- basis of most of the techniques used to assign functions to unknown genes
Module 5
Gene Inactivation
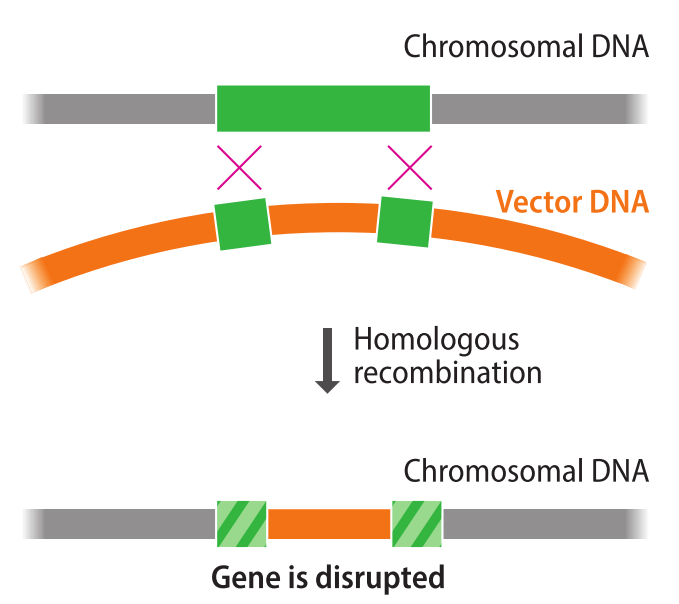
The easiest way to inactivate a specific gene is to disrupt it with an unrelated segment of DNA. This can be achieved by _____ _____. The vector carries two segments of DNA that match the _____ of the gene to be inactivated. These end segments _____ with the chromosomal copy of the target gene. As a result, the target gene becomes disrupted.
- homologous recombination
- ends
- recombine
Module 5
Gene Inactivation w/Homologous Recombination
Deletion Cassette
- deletion cassette consists of
- promoter sequence
- followed by an antibiotic-resistance gene
- new segments of DNA are attached to either end
- they have identical parts of the yeast gene to be inactivated
- flanked by two restriction sites
- start and end segments of the target gene are inserted into the restriction sites, and the vector is introduced into yeast cells
- Recombination between the gene segments in the vector and the chromosomal copy of the target gene results in disruption of the latter
- Cells in which the disruption has occurred are identifiable because they now express the antibiotic-resistance gene and will grow agar medium containing geneticin
Module 5
Gene Inactivation w/Homologous Recombination
barcode deletion
- A high-throughput version of the gene inactivation method
- uses a modified version of the basic deletion cassette
- also includes two 20-nucleotide barcode sequences
- different for each deletion, that act as tags for that particular mutant
- Each barcode is flanked by the same pair of sequences and so can be amplified by a single PCR
- groups of mutated yeast strains, each with a different inactivated gene, can be mixed together and their phenotypes can be screened in a single experiment
- the relative abundance of each barcode indicates the abundance of each mutant after growth in glucoserich medium
- Barcodes that are absent or present only at low abundance indicate mutants whose inactivated genes were needed for growth under these conditions.
Module 5
Gene Inactivation w/Homologous Recombination
model organism, knockout
- engineered embryonic stem cell is injected into a mouse embryo
- mouse embryo develops and gives rise to a chimera (mouse whose cells are a mixture of mutant ones), derived from the engineered ES cells embryonic site and nonmutant ones, derived from all the other cells in the embryo
- This is still not quite what we want, so the chimeric mice are allowed to mate with one another
- Some of the offspring result from fusion of two mutant gametes producing knockout mice
- works well for many gene inactivations, but some are lethal and so cannot be studied in a homozygous knockout mouse
- Instead, a heterozygous mouse is obtained in the hope that the phenotypic effect of the gene inactivation will be apparent even though the mouse still has one correct copy of the gene being studied
Module 5
Gene Inactivation w/out Homologous Recombination
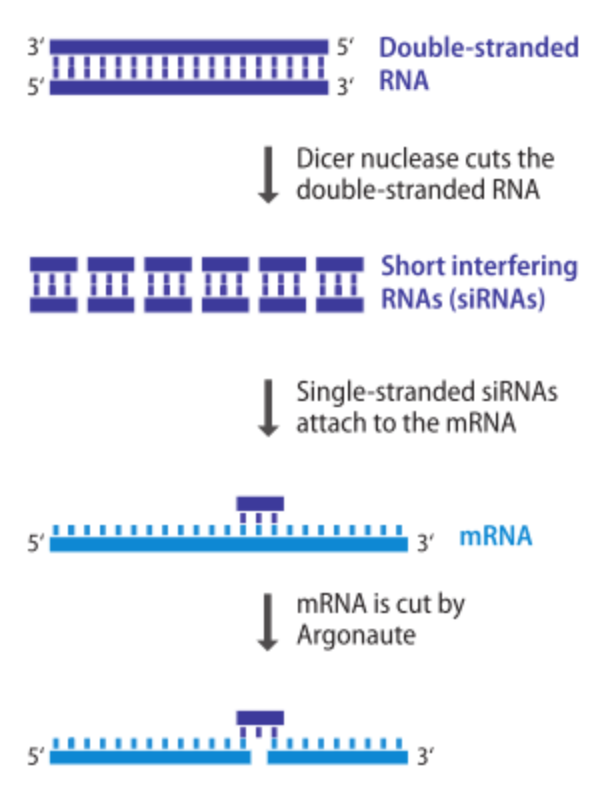
RNA interference, or RNAi
- natural processes by which short RNA molecules influence gene expression in living cells
- provides a means of silencing the expression of a target gene
- doesn’t disrupt gene itself but destroys its mRNA
- short double-stranded RNA molecules, whose sequences match that of the mRNA being targeted are introduced into the cell
- double-stranded RNAs are broken down into shorter molecules, which induce degradation of the mRNA
Module 5
Gene Inactivation w/out Homologous Recombination
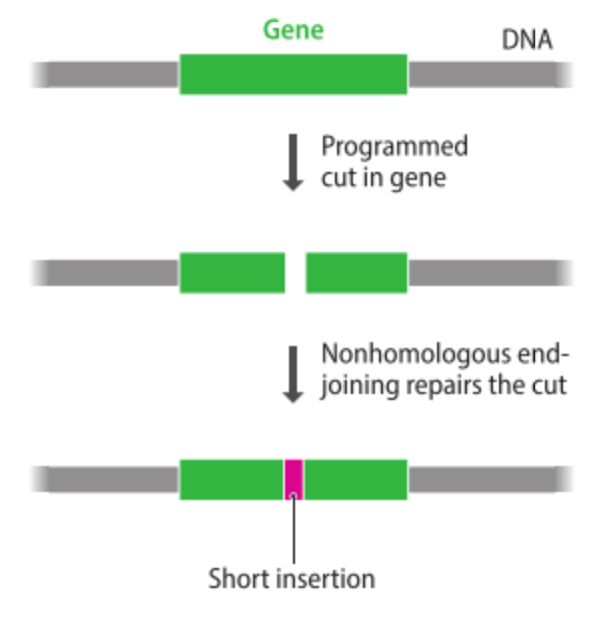
programmable nuclease
- a nuclease that can be directed to a specific site in a genome
- can be programmed to make a double-stranded cut in a selected gene
- stimulates nonhomologous end-joining
- results in a short insertion or deletion which will inactivate the gene
- creates a true knockout
- example: Cas9 endonuclease
Module 5
Gene Inactivation w/out Homologous Recombination
RNA interference, or RNAi issues
- does not always result in complete silencing of the target gene
- Often the silencing is incomplete and is referred to as knockdown rather than knockout
- so short that offtarget effects are possible
- interfering RNAs bind to mRNAs other than the targets, resulting in silencing of more than one gene
- In mammals it often results in activation of signaling proteins called interferons, which stimulate an antiviral defense resulting in phenotypic changes that can mask the specific change occurring due to silencing of the target gene.
Module 5
Gene overexpression
- used to assess function
- organism is engineered w/test gene much more active than normal (gain of function) to determine what changes, if any, this has on the phenotype
- a multicopy vector is used, that multiplies inside the host organism to 40–200 copies per cell and also contain a highly active promoter sequence
- must be treated w/caution because gene product may synthesized in excessive amounts, possibly in tissues in which the gene is normally inactive making it difficult to distinguish a phenotype change that is due to the specific function of an overexpressed gene
Module 5
The critical aspect of a gene inactivation or overexpression experiment is the need to identify a phenotypic change. This can be much more difficult than it sounds. Why?
- the range of phenotypes that must be examined is immense
- effect of gene inactivation can be very subtle and may not be recognized
- many gene inactivations appear to give no discernible phenotypic change
- effects of these genes only become apparent, if at all, when the cells are grown under a range of different conditions or when groups of genes that contribute to the same phenotype are co-inactivated
- In the human genome, there appears to be a subset of several hundred genes that are nonessential, both copies of which can be inactivated, due to natural mutation, without any discernible effect on the health of the individual.
- These observations suggest that a complete functional annotation of the genomes of many species will not be achievable by approaches that are based solely on gene inactivation or overexpression.






















