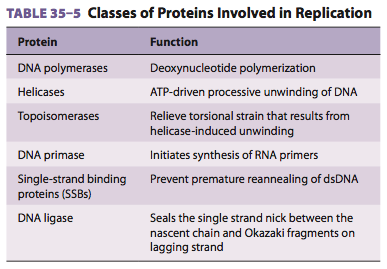
Which types of proteins are involved in DNA synthesis and what is their function?
What are the main differences btw pro- and eukaryotic DNA replication?
in eukaryotes:
- many ORC (instead of oriC) b/c much lower rate of replication → replication bubbles instead of replication fork
- SSBs are called replication protein A (RPA)
- different DNA polymerases
- DNA has to be rearranged into chromosomes after replication
Compare speed and error rate (fidelity) of DNA replication (in pro-/eukaryotes), RNA transcription and protein translation.
- DNA replication: very low error rate (high fidelity) ~ 10-8 - 10-9, a bit slower in eukaryotes due to nucleosomal interactions
- RNA transcription/translation: slower than replication, much higher error rate ~ 10-4
Differentiate btw prokaryotic DNA polymerases.
Which ones are involved in DNA replication?
- Pol I: DNA replication (RNA primer removal), DNA repair
- Pol II: DNA repair
- Pol III: DNA replication
Which factors contribute to the high fidelity of DNA replication in prokaryotes?
- intrinsic fidelity of DNA polymerase: 10-3 - -4
sensing dNTP complementarity to template - exonuclease proofreading of DNA polymerase: 10-2 - -3
sensing primer complementarity to template - mismatch repair: 10-2
sensing complementarity of 2 strands, distinguishing parental/daughter strands
⇒ overall: 10-8 - 10-9
Differentiate btw eukaryotic DNA polymerases.
Which ones are involved in DNA replication?
- Pol α: DNA replication (primer synthesis) = primase
- Pol β: base excision repair
- Pol γ: mitochondrial DNA replication/repair
- Pol δ: DNA replication (lagging strand synthesis), nucleotide/base excision repair
- Pol ε: DNA replication (leading strand synthesis), nucleotide/base excision repair
Which reaction is catalyzed by DNA polymerases (in eukaryotes and prokaryotes)?
dNTP + OH- on 3’ end as primer
→ DNA polymer w/ phosphodiester bonds + PPi
BUT: require template strand + Mg2+ as cofactor
⇒ synthesizes 3’→5’, daughter strand formed 5’→3’
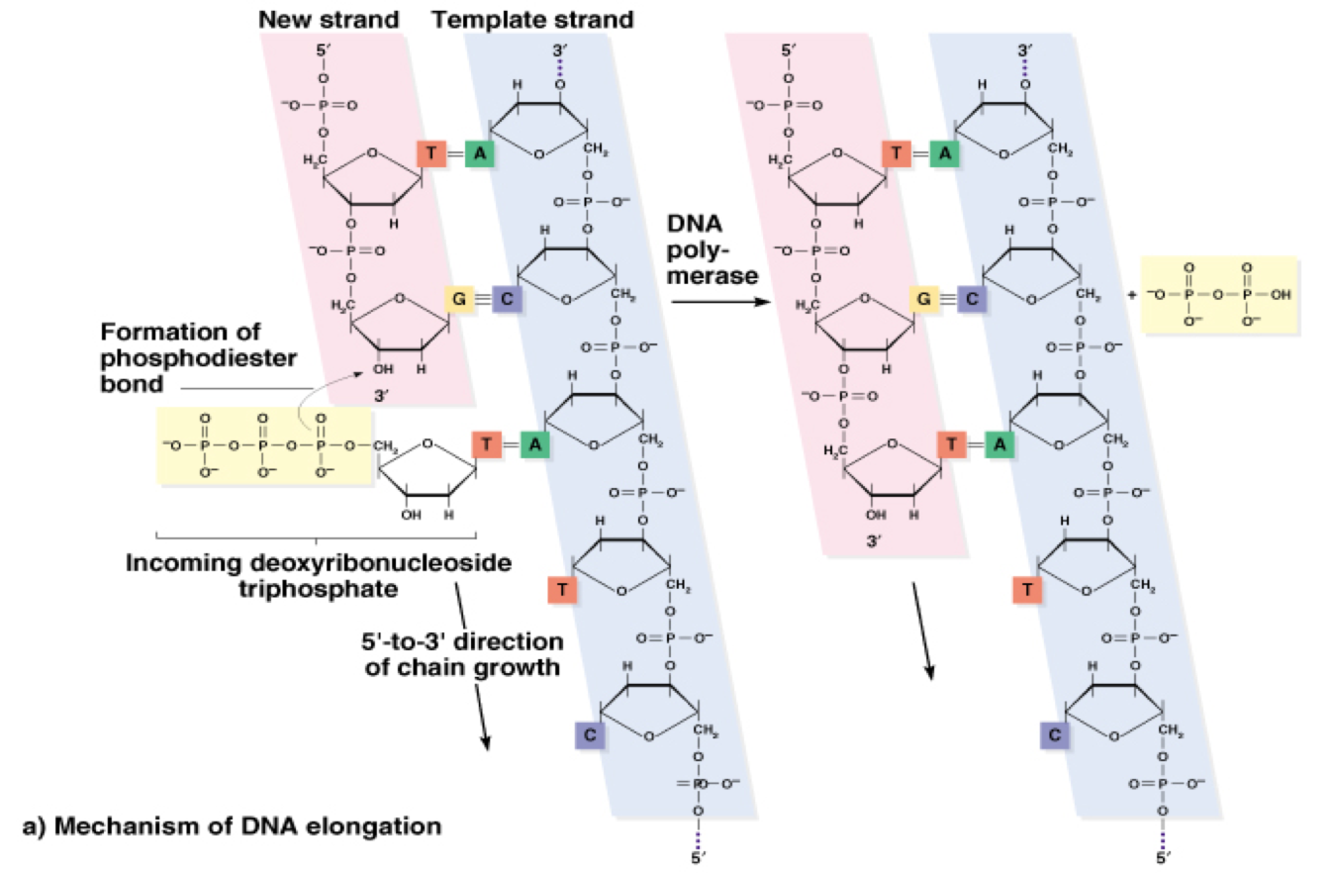
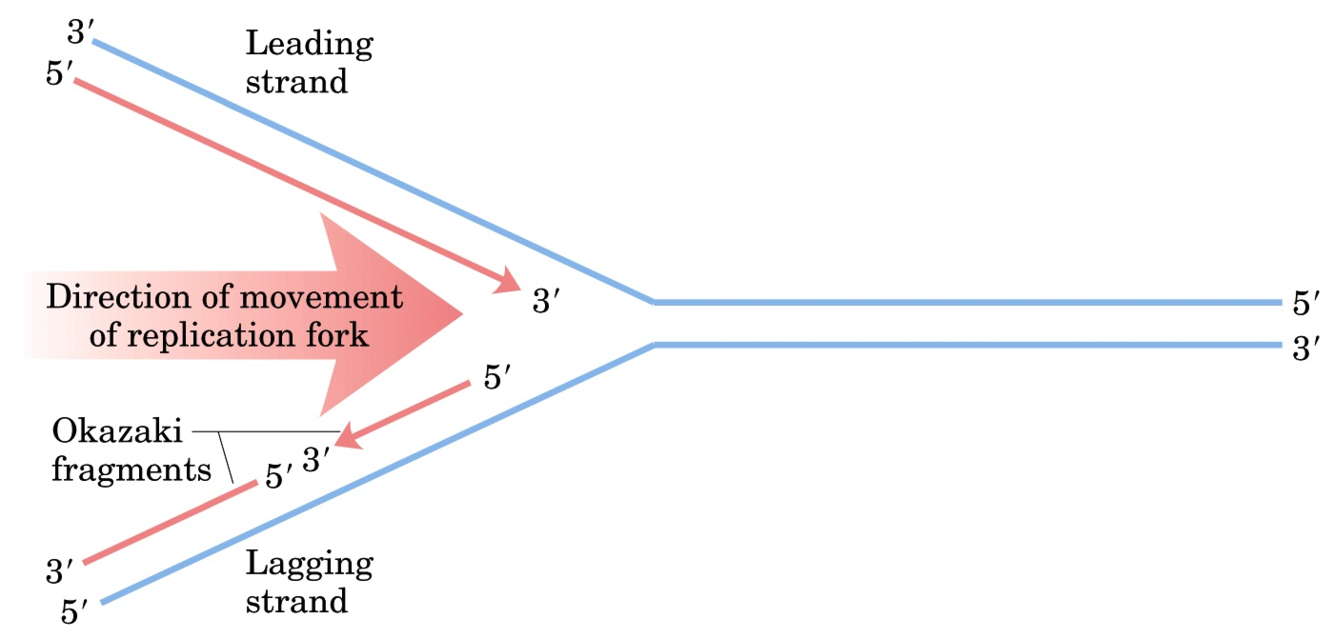
How are both DNA strands synthesized?
- leading strand synthesized continuously
- lagging strand synthesized discontinously, Okazaki fragments are formed (in prokaryotes ~ 1000 bps, in eukaryotes ~ 200 bps)
⇒ bidirectional synthesis in replication bubble
How is mismatch basepairing recognized by DNA polymerases?
DNA polymerase recognizes differences in atom distances + bond angles
What is special abt DNA pol I?
- coded by pol A: mutants lack proofreading activity
- monomer
only prokaryotic DNA pol w/ 3 activities b/c involved in replication and repair
- 3’→5’ polymerase activity
- 5’→3’ exonuclease (essential for initial removal of RNA primer)
- 3’→5’ exonuclease (proofreading)
How does DNA pol I know when to use its polymerase or proofreading activity?
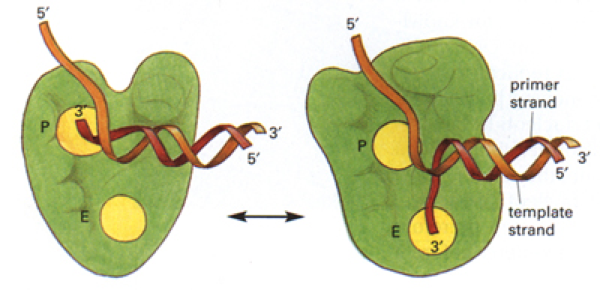
Klenow segment of DNA pol I has P and E site
- when template connected to primer passing through P site: polymerisation
- when template unwinds from primer + translocates to pass through E site: editing
What is the function of DNA pol II?
participates in repair, requires duplex template and primer
- coded by pol B: mutation is not lethal
- multimeric (4+)
only 3’→5’ exonuclease activity (proofreading)
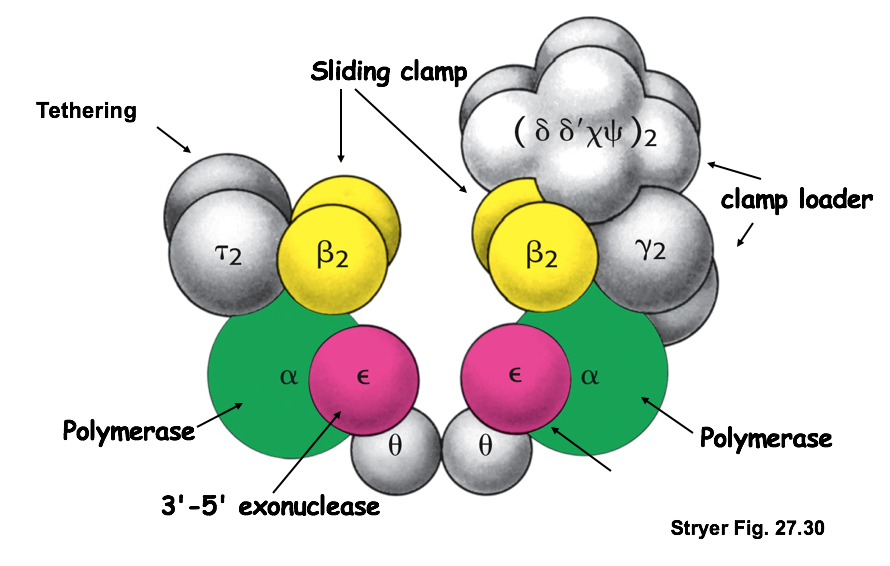
Describe the function of the structural elements of DNA pol III.
2 DNA pol III form an asymmetric dimer w/ multiple subunits
- each α subunit: 3’→5’ polymerase activity
- β subunits: act as sliding clamp, keep enzyme attached to DNA
- γ subunit: acts as clamp loader for lagging strand Okazaki fragment, help β subunits to form unit + bind to DNA
- each ε subunit: 3’→5’ exonuclease activity (proofreading stimulated by Θ subunit)
- τ subunits: dimerize, tether both enzymes together
What is the function of DNA pol III?
replicates prokaryotic DNA
- coded by pol C: mutants are temperature sensitive, conditionally lethal
3’→5’ polymerase activity +
3’→5’ exonuclease activity (proofreading)
What is the mechanism of 5’→3’ exonuclease activity of DNA pol I?
removes RNA primer on lagging strand + fills in the necessary nucleotides between the Okazaki fragments in 5’→3’ direction, proofreading for mistakes as it goes
What is the first step of the initiation of DNA replication in prokaryotes?
initiatior protein DnaA activated by ATP binding, multiple DnaAs bind to AT-rich 4bp regions of oriC (2 H-bonds, easier to unzip)
→ local denaturation + unwinding
What happens during replication in prokaryotes once the dsDNA is partially unwound?
recruitment of DnaB (5’→3 helicase), binds to lagging strand to start its action at AT-rich 13 bp regions of oriC = DUE (DNA unwinding elements)
→ ATP hydrolysis to unwind DNA to ssDNA
NOTE: binds to DnaG during elongation
The DNA is now “fully” unwound to ssDNA.
What happens now in prokaryotes?
(still part of initiation)
recruitment of DNA gyrase (topoisomerase I)
→ relieves the torsional stress caused by unwinding the double helix
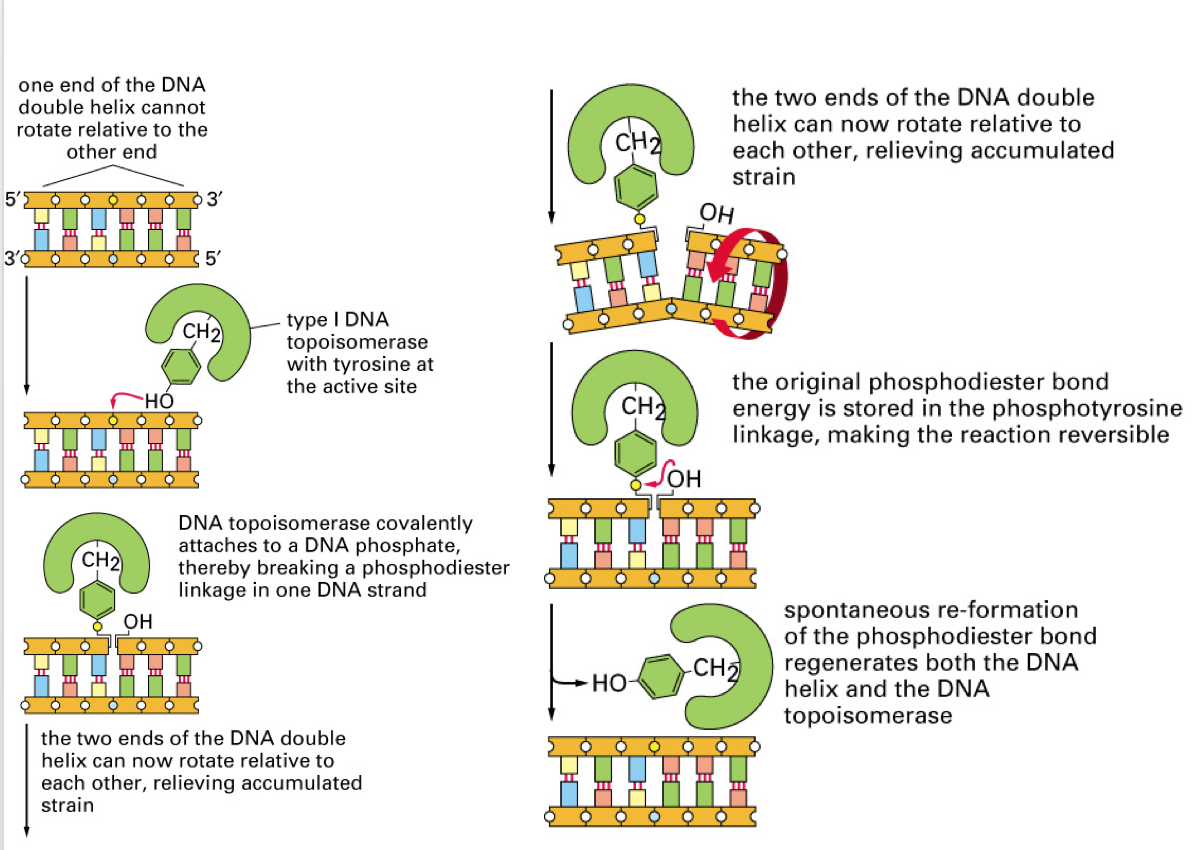
Describe the mechanism of topoisomerase I
NOTE: topoisomerase I causes single strand break
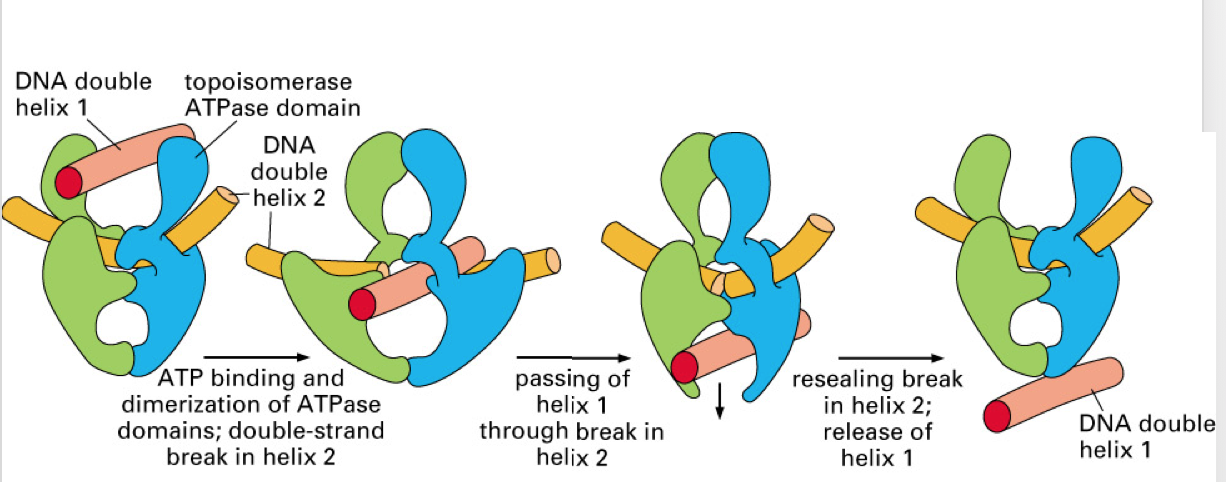
Describe the mechanism of topoisomerase II.
NOTE: topoisomerase II causes double strand break
List therapeutic implications related to topoisomerases.
- antibacterial quinolones (ex: ciprofloxacin) also targets bacterial topoisomerase I (DNA gyrase)
- topoisomerase I and II poisons used in chemotherapy
- topo I: topotectan
- topo II: daunorubicin, doxorubicin, etoposide
How is the replication fork kept open after unwinding in prokaryotes?
SSBs (single-strand binding proteins) bind to lagging strand + stabilize ssDNAs to maintain the replication bubble
Once the two strands are seperated, elongation begins…
What has to happen first though? In prokaryotes.
DnaG (primase) binds to leading and lagging strand to synthesize RNA primers to the template strands
- leading strand receives 1 RNA primer
- lagging strand receives several primers
NOTE: binds here to DnaB (helicase) to form primosome
What happens now, after attachment of primers to both DNA strands? In prokaryotes.
DNA pol III dimer binds w/ β subunits (sliding clamp) to leading strand, w/ γ subunits (clamp loader) to lagging strand
→ synthesizes daughter strands, beginning at 3’-OH of primer/s
- leading strand extended continuously
- lagging strand extended discontinuously forming Okazaki fragments
NOTE: τ subunits tether both dimers together, so that lagging strand synthesizing DNA pol III doesn’t leave replication bubble




