What is the rationale of using assays which test for known mutations?
Many applications require the ability to distinguish between two alleles that may differ only by a single nucleotide e.g. in diagnostics to distinguish between a disease-causing point mutation and a normal allele.
Genotyping by Sanger sequencing is not always efficient/economical when testing for a known sequence variant; there is no need to sequence a whole gene/exon, just a targeted region of interest.
What scenarios might methods for screening for a specific sequence variant become useful?
- Diagnosis of disease with limited allelic heterogeneity
- Detection of common pathogenic sequence variants
- Diagnosis within a family i.e. testing family members once the mutation has been characterised
- Testing ‘normal’ controls to determine whether a candidate pathogenic sequence variant is actually a polymorphism
- SNP genotyping
List 6 commonly used techniques for known mutation detection.
- Allele-specific PCR amplification
- Amplification refractory mutation system (ARMS)
- Minisequencing
- Oligonucleotide Ligation assay (OLA)
- Pyrosequencing
- Restriction Enzyme Digest
Describe the Allele-specific PCR amplification technique
- A common reverse primer and two forward allele-specific primers amplify two allele-specific PCR products of different lengths
- separated by agarose gel electrophoresis.
- This techniquemakes use of the fact that under suitable conditions amplification will not take place when the 3’ end nucleotide is not perfectly base paired, thereby distinguishing the two alleles.
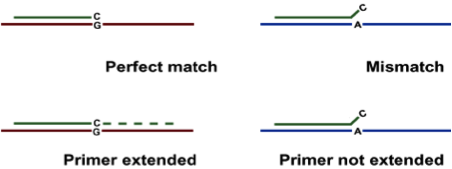
Describe the ARMS PCR method.
ARMS is an example of allele specific PCR
- Paired PCR reactions are performed in two separate tubes, one primer is the same in both reactions and the other is specific for either the normal or mutant sequence.
- The allele specific primers differ in their 3’ end nucleotides, designed to base pair with the variable nucleotide that distinguishes the alleles. Amplification does not occur when the 3’ end nucleotide does not match, allowing the two alleles to be distinguished.
- The location of the common primer can be designed so that different product sizes are given for different mutations and the reactions can be multiplexed and separated by gel electrophoresis.
- Pairs of mutation specific primers can be given different fluorescent labels or 5’ extensions of different sizes to be able to distinguish between PCR products (by gel electrophoresis or capillary fluorescent electrophoresis).

Whilst no longer in common use give an example of a commercially available ARMS-PCR assay.
- Elucigene CF29 kit for Cystic Fibrosis common mutations - CFTR known mutation detection
- Thrombosis risk panel (including the common risk factor polymorphisms Factor V Leiden R506Q, Prothrombin 20210A and MTHFR 677C)
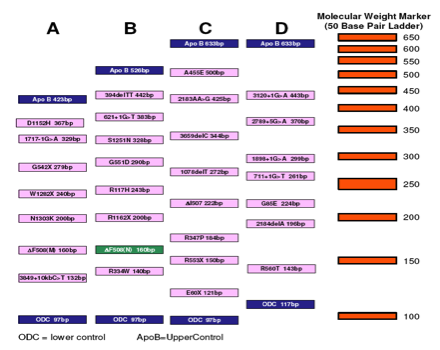
Give some advantages of ARMS-PCR.
- Quick, sensitive, simple
- -Once designed and optimised is relatively inexpensive
- -Does not require specialised equipment
- -Multiplexing is possible
Give some disadvantages of ARMS-PCR.
- Cannot detect unknown mutations
- Polymorphisms under the primer site can affect primer binding
- Designing assays and multiplexing can be difficult so commercial kits are often required, with kits heterozygosity may need confirmation with another method
- Cross reactivity may pick up the wrong mutation e.g. CF p.Phe508del/p.Phe508Cys
- Prone to contamination e.g. MCC in prenatal samples, and methods for detecting MCC may not be as sensitive as the ARMS assay
- A single primer mismatch is often not sufficient to prevent bleed through and a second mismatch is usually incorporated (generally if a 3’ mismatch is weak a strong secondary mismatch is engineered, if it is strong a weak secondary is introduced)
Breifly describe the minisequencing method.
- The minisequencing method is based on the incorporation of a single fluorescently labelled ddNTP at the 3’ end of a special oligonuclotide which is complementary to the sequence and is located one nucleotide before the examined polymorphic site
- The incorporation of one of four ddNTPs labelled by differently collared dyes depends on the genotype.
- After reaction, the products of the reaction are separated by capillary electrophoresis
- The colour of the single signal (hom) or two signals (het) allows the identification of the incorporated ddNTP and thus the complementary nucleotide in the DNA sequence
- Furthermore, If multi length oligonucleotides are used, several different polymorphisms/mutations in one multiplex reaction can be detected
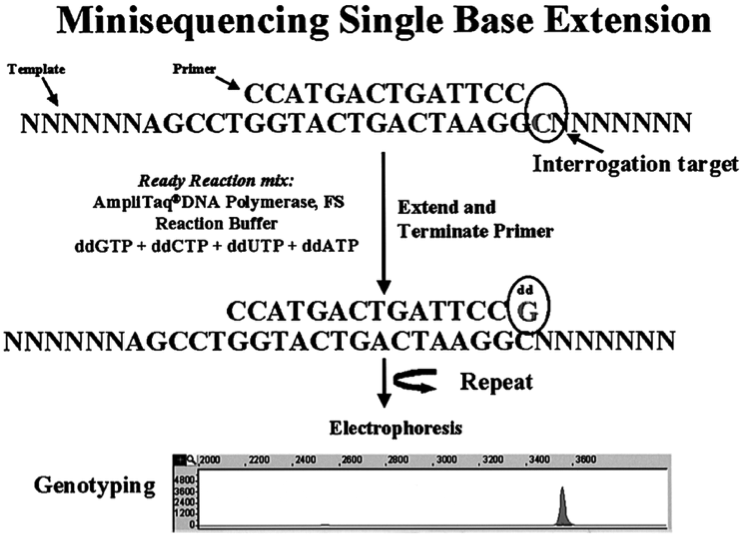
Briefly describe the OLA method
OLA consists of two stages, a PCR amplification and an OLA in a single tube format.
- PCR primer is hybridised to the target sequence. The primers are designed with either normal or mutant nucleotide(s) at the 3’end and a tail of different lengths to distinguish PCR products based on size. A PCR reaction is performed.
- A common primer contains a fluorescent dye marker (e.g. FAM) at the 3’end and meets the first primer over the nucleotide position that is altered in the mutant allele. If the 3’end of the first primer matches perfectly with the target DNA, both the primers will be ligated together. No ligation will occur when there is a mismatch between the 3’end of the first primer and the target DNA
- Two differently sized florescent fragments are produced discriminating the two alleles
- Reactions can be multiplexed by adding stuffer sequence of different lengths and different coloured dye labels in the oligonucleotides
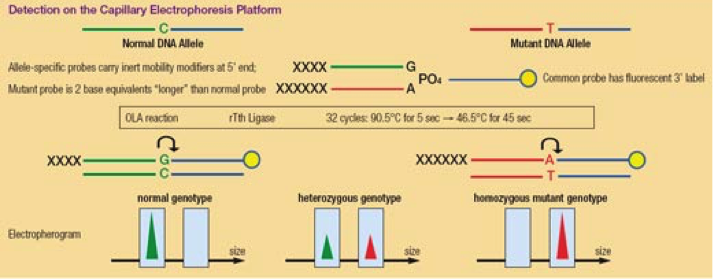
Give an examples of OLA in use in diagnostic laboratories.
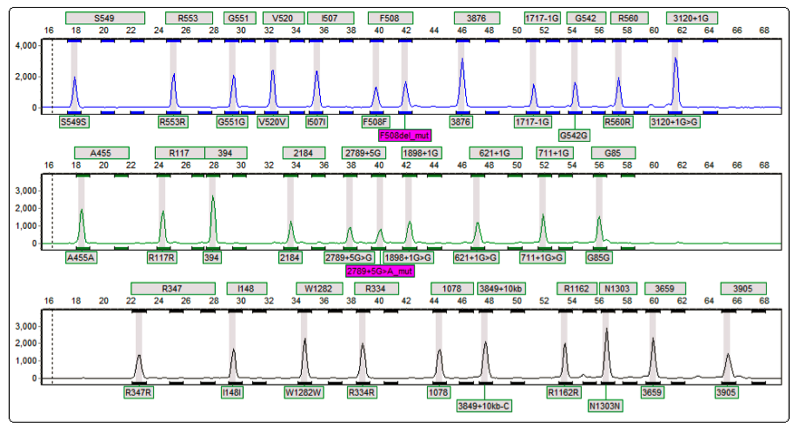
Abbot Molecular CFTR genotyping OLA assay: manufacture a commercial OLA kit, the Cystic Fibrosis Genotyping Assay for detecting common CFTR mutations
Image: A ligation product from the common p.Phe508del mutation is seen (purple box, top row), along with a product from the c.2789+5G>A mutation (purple box, middle row). This patient is a compound heterozygote.
Advantages of OLA
- Easy, quick, single tube assay
- -High throughput and automatable on a capillary sequencer
- -Can be multiplexed
- -Can be automated
Disadvantages of OLA
- Requires an automated sequencer
- -In-house design is difficult and expensive (although customisable products are available e.g. SNPlex from Applied Biosystems
- -Fairly reliant on commercial kits, kits can be expensive
- -Cannot detect rare mutations
- -Polymorphisms can affect primer binding site
What is pyrosequencing and what applications can it be used for?
- A method of examining very short sequences of DNA (1-5 nucleotides) adjacent to a defined starting point. Can detect any point mutation as long as the expected sequence and position of the mutation are known.
- The main use is for SNP genotyping, in which only one or two bases are sequenced.
How does pyrosequencing work?
- When a dNTP is added to an open 3’ DNA strand pyrophosphate is released. A cocktail of enzymes is used in pyrosequencing which couples this pyrophosphate to light emission by luciferase.
- A primer is hybridised to the test DNA and offered each dNTP in turn.
- When the correct dNTP is present the primer can be extended and a flash of light is emitted and recorded. The amount of light emitted is proportional to the number of bases incorporated.
- Any bases not incorporated are degraded by apyrase and the reaction can start with another nucleotide.
Name an examples of how pyrosequencing used in diagnostic laboratories.
- Pyrosequencing is quantitative so is useful for somatic mutation detection in cancer e.g. analysis of tumour tissue for: p.(Val600Glu)(6) in BRAF; codons 12, 13, 61 and 146 in KRAS(7); codons 12, 13, 59 and 61 of NRAS.
- Common mutations associated Parkinsons Disease in LRRK2
- HFE mutations H63D and S65C and C282Y
The c.1849G>T, encoding p.(Val617Phe), mutation of JAK2 (a tyrosine kinase gene) is widespread in chronic myeloproliferative disorders.
The sensitivity of the Pyrosequencing JAK2 assay enables accurate quantification of the mutation in minor subpopulations of blood/bone marrow myeloid cells.
Can you interpret the trace seen in the image?
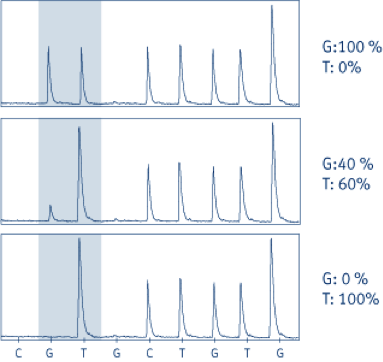
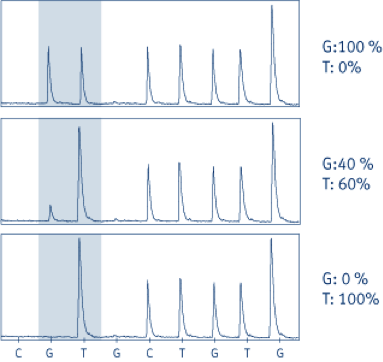
Pyrogram® traces of JAK2 analyses. The sequence read is (G/T)TCTGTGG. The shaded area indicates the mutation site. The G peak in the shaded box varies between 0 and 1 (normalized units), whereas the T peak varies between 1 and 2, since the subsequent nucleotide in the sequence is T.
Top box: Normal control
Middle box: Philadelphia chromosome negative CML, normal G is 40% and mutant allele is 60%.
Bottom box: HEL cell line showing 100% mutant T allele.
ADvantages of pyrosequencing
- -Will detect any sequence change if the reference sequence is known
- -Useful for analysis of many types of genetic change e.g. SNPs, RNA allelic imbalances, DNA methylation & gene copy number
- -Rapid analysis time
- -High throughput
- -Cheap if high throughput
- -Sensitive (can detect low level mutations down to a level of 2%) and quantitative
- -Works well with degraded DNA e.g. from tumour samples
Disavantages of pyrosequencing
- -Specialist automated sequencer is required
- -Lots of PCR cycles required to use all primer increasing the risk of contamination
- -Read lengths are short so need separate reactions for different mutation sites
- -Secondary structure can be a big problem
- -Cannot sequence long homopolymer tracts as the light emission signal is not completely linear for each base added
- -“Unexpected” sequence variants need confirming with sequencing
- -Sensitivity of mutation detection is reduced when no new peak in the Pyrogram is created e.g. GGT > GTT
What are Restriction Enzymes and how can they be used for known mutation detection?
- Restriction enzymes are enzymes that make double stranded breaks in DNA at specific recognition sites
- Can be used when a base substitution creates or abolishes a recognition site of a restriction endonuclease
- However for diagnostic testing the sequence variant needs to create a restriction site
Give an overview of the method for restriction enzyme digests.
- A suitable length sequence containing the potential mutation is amplified, the PCR product is digested with the relevant restriction enzyme and the products separated by gel electrophoresis to see if a cut has occurred or not.
- It is good practice to design a PCR containing a second site for the selected restriction enzyme. This second site will be present regardless of whether the sequence variant under investigation is present so it will act as a positive control for digestion and confirm enzyme activity.
How can restriction enzyme disgests be modified to further broaden their utility?
- Variations include methylation specific restriction digests in which the enzyme is methylation sensitive and will only digest methylated DNA.
- Even when a mutation does not occur within a known recognition sequence it may be possible to introduce a site by PCR mutagenesis using carefully designed primers
Give an example of utilising PCR mutagenesis and restriction enzyme digests to confirm the the presence of a mutation.
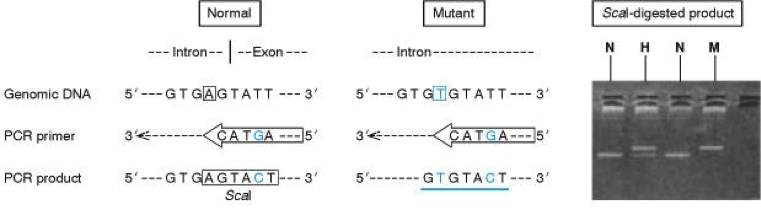
An A>T mutation in the intron 4 splice site of the FACC gene does not create or abolish a restriction site.
The PCR primer stops short of this altered base, but has a single base mismatch (blue G) in a noncritical position which does not prevent it hybridizing to and amplifying both the normal and mutant sequences.
The mismatch in the primer introduces an AGTACT restriction site for ScaI into the PCR product from the normal sequence. The ScaI-digested product from homozygous normal (N), heterozygous (H) and homozygous mutant (M) patients is shown.
Advantages of restriction enzyme digests?
- Cheap, simple, easy, quick
- No specialist equipment required


