What is “Third Generation Sequencing”?
Third Generation Sequencing is a class of DNA sequencing that works by reading very long fragments of nucleotide sequences at the single molecule level.
In contrast to NGS that requires breaking long strands of DNA into small segments and clonal amplification prior sequencing.
What are the proposed advantages of SMS over NGS?
- Small amount of starting material
- Simplify template preparation and reduce amount of template required
- Higher throughput – unlike 2nd generation, in TGS many hundreds to millions of sequencing reactions can be carried out asynchronously.
- Lower projected cost per base – high fold coverage for less than $100 is now a reasonable goal
- Aim to sequence at a faster rate
- Longer read lengths
- Potentially less sensitive to GC content than 2nd generation sequencing
- Potential to detect epigenetic modifications such as methylation
- Redundant sequencing will result in higher consensus accuracy to enable rare variant detection and simplified data analysis
What are the key advantages to longer read lengths of SMS?
Longer read lengths will facilitate
- Enhanced de novo assembly
- haplotype detection
- chromosome phasing
- CNV detection
- recognition of insertions
- deletions and translocations
- detection of novel isoforms resulting from alternate splicing
- identification of chimeric transcripts.
- Sequencing of repetitive elements
What different chemistries have been developed for SMS?
- Sequencing by synthesis: Single molecule of DNA polymerase is observed as it synthesises a single molecule of DNA
- Nanopore: Single DNA molecule is threaded through a nanopore (biological or synthetic) and individual bases are detected as they pass through the nanopore.
- Synthetic approaches: Generating long read sequencing reads that rely on existing short-read technologies
What are the two different strategies to “Sequencing by synthesis”?
- Single molecule real time (SMRT) sequencing (developed by Pacific Biosciences “PacBio”)
- FRET sequencing (under development by Life Technology)
What is the principle of SMRT sequencing ?
- Based on monitoring polymerase activity while incorporating differently labelled nucleotides into the DNA strand.
- SMRT technology utilises zero-mode waveguide (ZMW) – a hole measuring 30-70 nm in diameter.
- A single DNA polymerase is anchored to the bottom surface of each ZMW
- Each nucleotide carries a base-specific fluorescent label which is released when being incorporated by the polymerase.
- All dNTPs are present at the same time and dNTP incorporation is continuously visualised with a laser
- the enzyme holds each nucleotide within the detection volume for very short time period but is enough to engage the fluorophore to emits fluorescence.
- The template strand move forward one bp and the cycle continues.
What are the advantages of SMRT sequencing?
- very fast – DNA polymerase continues to incorporate bases at a speed of tens per second = kilobases of DNA in minutes
- can discriminate between methylated and unmethylated bases which could have clinical utility
- Can accurately sequence repetative regions
- Very low error rate after repeated sequencing of template
What are the disadvantages of SMRT sequencing?
- Limited throughput: Limited by the number of ZMWs that can be read per SMRT cell
- PacBio have recently launched the sequel system – reportedly has 7X throughput of RS II.
- Expensive ~$1000/Gb = >$3000 per himan genome = out of reach of small labs.
What is the underlying principle of “nanopore” single molecule sequencing?
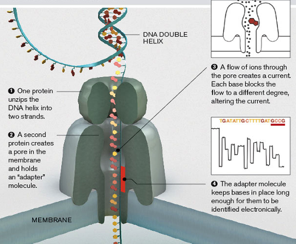
- Underlying principle is that a ssDNA molecule is electrophoretically driven through a nanoscale pore in a strict linear sequence.
- A nanopore is a very small hole through a thin membrane submerged in salt buffer solution.
- When an electrical potential is applied to solutions on either side of the pore, it drives dissolved salt ions through the pore, establishing electrical current.
- Molecules passing through the pore can measurably change the ionic current
- The amplitude and duration of these transient current blockades is determined by the physical and chemical properties of the target molecules
What are the two classess of nanopore chemistry?
- Biological nanopores: Transmembrane protein channels inserted into a substrate, eg planar lipid bilayers, liposomes or polymer films. They have a well-defined and highly reproducible nanopore size and structure, and can be modified easily using molecular biology techniques
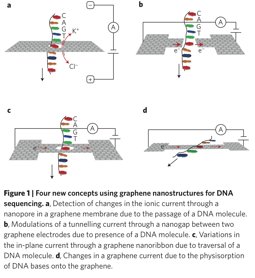
- Graphene Nanopore: Graphene is made up of layers of graphite (1nm thick) with drilled pores (from 0.3 nm in diameter - the distance between two DNA bases is 0.5nm). At one atom thick, graphene is believed to be the thinnest membrane able to separate two liquid compartments
What characyeristic are important for selecting a biological nanopore?
- Temperature and pH stability: a-hemolysin is stable at 100°C within a wide pH range.
- Size of channel: a-hemolysin channel is 1.4nm at its narrowest point so ssDNA, which has a diameter of 1-1.3nm, can pass through the pore
What are the limitations to using biological nanopores?
- Due to the length of the pore the shift in voltage due to the presence of a DNA molecule is caused by the string of bases within the pore rather than by an individual nucleotide.
- This string is called a k-mer (k=length of DNA string).
- Therefore rather than having 1-4 possible signals there are more than 1000 – one for each k-mer.
- This is increased if modified bases are taken into account.
- This results in a large error rate of up to 30%, particularly with indels, and homopolymers longer than the k-mer length can be difficult to identify.
- The rapid DNA translocation velocity through the pore (1-3ms/base) limits the identification of single nucleotides, and there is a requirement for ultra-precise high-speed DNA detection systems and modified enzymes to slow down translocation.
What key advantages of nanopore sequencing?
- Potential to be very quick and very cheap compared to NGS
- OGT claim that their instruments are set up so that the sequencing data can be analysed in real time as the reaction is happening
- Therefore the user can access whether enough data/coverage has been obtained and choose when the experiment is complete rather than hoping the coverage per sample is sufficient from a given instrument run.
Who is the market leading nanopore SMS provider and what are they key instruments?
- Oxford Nanopore is the market leader of nanopore SMS.
- MinION nanopore sequencing was used to genotype Salmonella strains from an outbreak in an English hospital within 6 hours (Quick et al 2015), and to sequence Ebola strains in the field and monitor transmission history and evolution of the virus as the outbreak unfolded (Quick et al 2016).
- PromethION: Recently developed. Ultra-high throughput platform reported to include 48 individual flowcells each with 3000 pores running at 500bp/sec. Works out to ~2-4 terabytes for a 2 day run. Cost and error profile not available yet.
What are the advantages and disadvantages of using Graphine solid state pores over biological pores for SMS?
Advantages
- Solid-state nanopores are more stable than biological nanopores
- There is more control over pore diameter and channel length
- lower sensitivity to external parameters such as pH, temperature, salt concentration, and mechanical stress
- Well suited for massive upscaling and device integration on chip
Disadvantages
- Lack of true atomic control of pore = high translocation rates
- Increased noise levels
What is the principle behind “Synthetic approaches” to generating long read sequences from single molecules?
- DNA is fragmented into large segments of ~10kb
- Then partitioned into either microtitre wells or an emulsion such that one molecule exists in each partition.
- The individual molecules are sheared into smaller pieces and barcoded, then sequenced on existing NGS instruments.
- Fragments with the same barcode have therefore been generated from the same original large fragment and the sequence can be reassembled.
Who are the two main provider of synthetic long read products?
- Illumina Synthetic Long-Read (formally Moleculo)
- 10X Genomics (Chromium)
What are the advantages and disadvantages of using synthetic long-read?
Pro’s
- Enables utilisation of existing NGS hardward (no capital investment)
- Can use the single-molecule data to phase variants (prove in cis/trans) and detect large copy number changes (dels/dups)
Con’s
- Because it relies on short-read sequencing chemistry it is still prone to errors in any region where the Illumina chemistry is biased, such as regions with high GC content or tandem repeats and flow-cell related errors.
- Obtaining sufficient coverage for de novo genome assembly can be expensive, often even greater than PacBio sequencing.
What is “Third Generation Mapping” and what is it’s application?
Single molecule technologies to determine the large-scale structural changes of DNA without sequencing.
Applications should be to replace microarrays and chromosome analysis which yield information about copy number and SVs but not SNVs.
What is the principle behind third generation mapping?
- Uses fluorescently tagged probes attached at “nicked” restriction digest site
- Each enzyme will generate a unique genome-wide ‘nicking’ fingerprint along DNA molecules.
- After imaging, the per-molecule fingerprints are assembled into larger optical maps, typically spanning many megabases of a chromosome.
- Maps can be compared to a sequence assembly to eference genome of how the sequences should be ordered and oriented along the chromosome to reveal structural changes, such as the rearrangement or fusion of two chromosomes.
Why have Third generation mapping methods not taken off?
- Existing methods are already very cheap and don’t require expensive hardware
- Third generation methods suffers from biases e.g. incomplete nicking of the DNA, causing a proportion of the digest sites remain unlabeled, and “fragile sites” where multiple nick sites in close proximity cause the DNA to systemically shear and limit the overall length of the map.


