What is ‘Mutation scanning’?
Mutation scanning is the search for novel sequence variants within a defined DNA fragment (without sequencing).
Some scanning methods do not identify the precise nature of the change to the DNA sequence, although some indicate the location of the mutation within the fragment analysed.
Scanning methods exploit different physical, chemical, and biological consequences of DNA sequence variation to indicate a change in the fragment being analysed.
Define the ideal mutation scanning method.
The ideal mutation scanning method would screen;
- kilobase lengths of DNA
- 100% sensitivity
- 100% specificity
- completely define the mutation
Previous common techniques are now largely superceeded by NGS and/or arrays.
You have to implement a mutation scanning test in your laboratory.
What factors must you consider before choosing which type of scanning method to select?
- Mutation Detection Sensitivity
- Suitability for Proposed Sample Type (e.g. Peripheral blood/tumour tissue)
- Suitability for Predicted Mutation Type (e.g. PTT detects only polypeptide-chain-terminating mutations)
- Features of the DNA Sequence Analyzed (e.g. Presence of polymorphisms)
- Health and Safety Considerations (e.g. Toxicity of chemicals used)
- Expected Requirements for Sample Throughput
- Capital Equipment Costs and Ongoing Running Costs
- Requirement for Post-PCR Manipulation (i.e. minimal number of post-PCR steps)
Name the three broad types of mutation scanning methods.
- Separation by physical differences (rely on differences in 3d conformation between SNPs)
- Seperation by chemical differences
- Enzymatic methods
Give three examples of mutation scanning methods which utilise ‘separation by physical differences’
- Single Strand Conformation Polymorphism (SSCP)
- Conformation Sensitive Gel Electrophoresis (CSGE)
- Denaturing High Performance Liquid Chromatography (DHPLC)
Give four examples of mutation scanning methods which utilise ‘seperation by chemical differences’
- High resolution melt curve analysis (HRM) (differences in melting temperatures between SNPs)
- Dynamic Allele Secific Hybridisation (SNP DASH) (differences in melting temperatures between SNPs)
- Mass spectrometry (MALDI-TOF/Sequenom) (differences in size and charge between SNPs)
- Protein truncation test (PTT) (only detects truncation mutations, relies on shorter than usual protein product)
Give five examples of mutation scanning methods which utilise ‘enzymatic methods ‘
- Chemical Cleavage Mismatch (CCM)
- Enzyme Cleavage of Mismatch (ECM)
- Base Excision Sequence Scanning (BESS)
- Cleavage Fragment Length Polymorphism (CFLP)
- Restriction Fragment Length Polymorphism (RFLP)
What is DHPLC?
- Denaturing High Performance Liquid Chromatography relies on the creation of hetero- and homo- duplexes of paired DNA in order to identify SNPs.
- The products are then separated using a slightly denaturing liquid chromatography column.
- The heteroduplexed positions are denatured and the products therefore move more slowly indicating that a SNP is presentin the fragment being analysed.
Briefly describe the MALDI-TOP mutation scanning method.
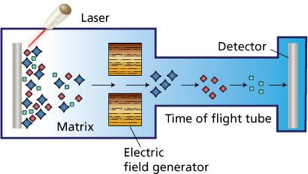
- Sample is placed in a UV-absorbing matrix pad and exposed to a short laser pulse.
- Ionised molecules are accelerated off the matrix pad (desorption) and move in an electric field towards the detector.
- The ‘time-of-flight’ required to reach the detector depends on the m/z of individual molecules.
Give 5 attributes of MALDI-TOF mutation scanning.
- Can measure DNA molecules up to 20 kD
- Accuracy of ±0.3%
- Can size oligonucleotides ~100 bases long faster than electrophoretic separation.
- Base composition can be established also.
- Can be used to analyse labelled proteins transcribed from the DNA to detect sequence variation or to directly analyse labelled DNA.
Give an example of a commercially available MALDI-TOF platform.
List the analytes that the platform can interrogate and give it’s genotyping throughput.
Sequenom MassARRAY platform
Used for analysis of;
- ffDNA to determine the fetal Rhesus D geneotype.
- methylation analysis (bisulphite treated DNA)
- panels of SNPs
- gene expression (mRNA analysis)
- DNA sequencing
Potential to multiplex up to 36plex, giving throughput of up to 138,000 genotypes/day.
Describe the method for utilising MALDI-TOF for genotyping of SNPs
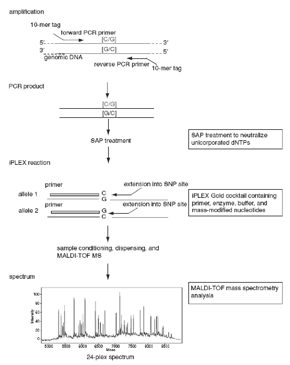
- PCR-based amplification of the region of interest.
- Locus-specific primer binds immediately upstream of the polymorphic site.
- Single nucleotide extension is performed using mass-modified (different mass for each base) dideoxynucleotide terminators.
- The mass of extended primer is used to determine the sequence at that particular site.
Describe the method for utilising MALDI-TOF for DNA sequencing
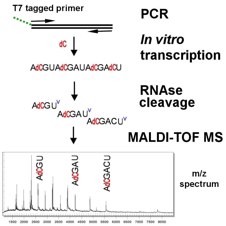
- PCR-derived DNA product undergoes in vitro transcription in four separate reactions (each with three rNTP bases and one specific dNTP).
- The use of dC prevents cleavage of C positions by RNase, which cleaves only after rU and produces three fragments of 4, 5 or 6 nucleotides (each with a characteristic m/z ratio).
- Analogous reactions occur for the other three bases.
- The four reactions are taken together and combined.
- Mass signal pattern obtain is compared with the expected m/z spectrum for the reference sequence under consideration.
- Any differences between experimental and reference DNA sequences will produce predictable shifts in the spectrum.
Name the advantages of MALDI-TOF
- Efficient ionisation of DNA molecules using MALDI techniques
- Accurate determination of mass – sufficient to allow base composition of DNA sequence from mass alone
- Faster than conventional methods e.g. electrophoretic-based methods
- High throughput – thousands of genotypes per day
- Allows allele frequencies of a point mutation to be estimated in analysis of a pooled sample
- Can be a relatively cheap method if the machine is already available in the laboratory and used for high throughput.
Name the disadvantages of MALDI-TOF
- Expensive
- Large, expensive equipment necessary
Briefly descibe the high resolution melt curve analysis (HRM) mutation scanning method.
- PCR-based amplification of region of interest in the presence of a specialised dsDNA binding dye which is highly-fluorescent only when bound to dsDNA.
- Fluorescence is measured whilst the amplicon is subjected to a gradual increase in temperature – when Tm is reached, DNA strands in the duplex are separated which releases the dye and results in a decrease in fluorescence.
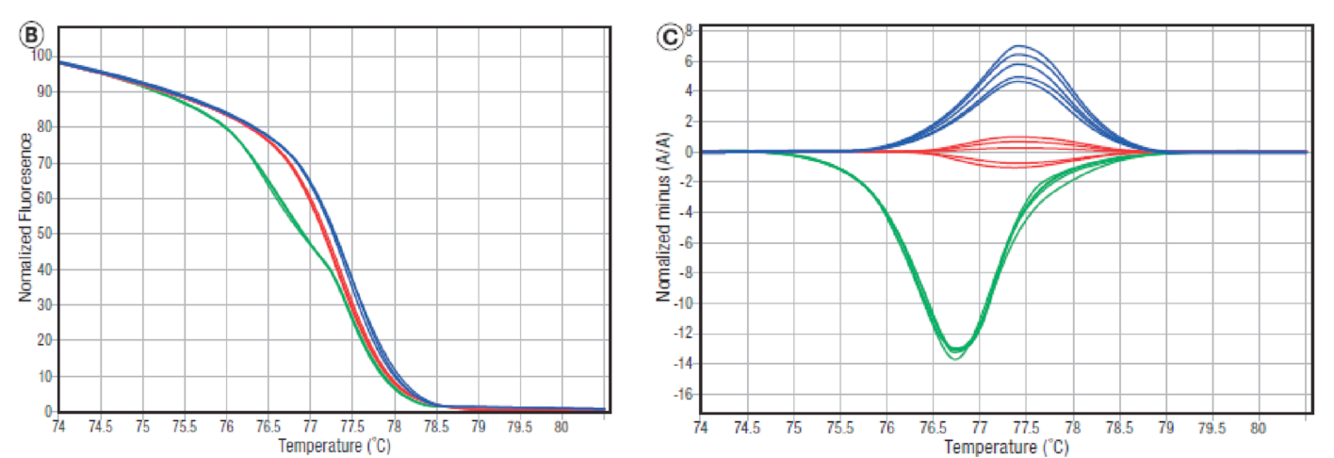
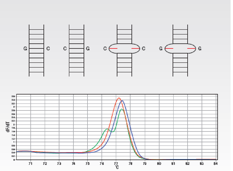
- Plot fluorescence vs. temperature. Raw data is shown on left below. Melting peaks are generated by plotting the first negative derivative of this data, shown on right below.
Intercalating Dyes are essential to the HRM scanning method.
What attributes should an Intercalating Dye have to be utilised in HRM.
- should not preferentially bind to any bases
- should not inhibit PCR and should not influence Tm.
Give examples of different types of Intercalating Dyes.
- Non-saturating dyes e.g. SYBR Green
Unsuitable for HRM! Stabilise duplex and inhibit polymerase. The released dye can also rebind to amplicon during melting.
- Saturating dyes e.g. SYT09
Can use at high concentration without altering reaction dynamics. Do not redistribute during melting as the amplicon is saturated.
- Release-on-demand dyes e.g. Evagreen
Added at non-saturating levels. Fluorescent signal is quenched when dye is in solution. Upon binding, quenching factor is released.
What other factors are critical for the success of HRM?
- The power of HRM depends upon the sensitivity of the instrument and the specific PCR reagents used.
- Certain instruments have been designed specifically for HRM analysis; rotary-based instruments should be used for high sensitivity and reproducibility.
- Whilst block-based instruments should be used for high-throughput and ease of handling.
How are the results of HRM interpreted?
- Following PCR, there will be a pool of 4 different amplicons for heterozygous samples as a composite of heterozygote (resulting from mispairing) and homozygote duplexes are created.
- Results in a mixture of four different melting profiles all combined into one fluorescence signal output.
- The melting profile of a heterozygote sample (in green) often displays a double peak compared to the single peak of the homozygotes.
- The relatively unstable mispaired heteroduplexes melt at a lower temperature than the correctly paired samples, so heterozygotes are easily identified.
What are the advantages of HRM mutation scanning?
- Quick! Melt curve analysis takes ~5 minutes following PCR
- Closed tube system – limits contamination.
- Cheap reagents (only high cost is the machine)
- High throughput.
- Can use PCR products for other applications following analysis e.g. sequencing if an aberrant melt profile observed.
What are the disadvantages of HRM mutation scanning?
- Not 100% sensitive – estimated that 93% of unique heterozygotes are distinguishable by HRM
- most, but not all, homozygotes can be distinguished by HRM
- ~4% of human SNPs are predicted to have the same Tm by nearest neighbour symmetry and so may be missed
- Not suitable for highly-polymorphic genes
- Need a small fragment (50-150bp) which contains no more than 1 or 2 ‘melt domains’
- Extensive optimisation – primer design, setting parameters, sensitivity depends on mutation etc.
- Best suited for analysing large number of samples to enable accurate data analysis
Describe the principle behind enzymatic screening methods.
- Utilises an endonuclease to cleave double stranded DNA at heteroduplexes
- Amplification of region of interest using PCR/RT-PCR.
- Digest with Surveyor enzyme.
- Either gel electrophoresis or analysis using WAVE system
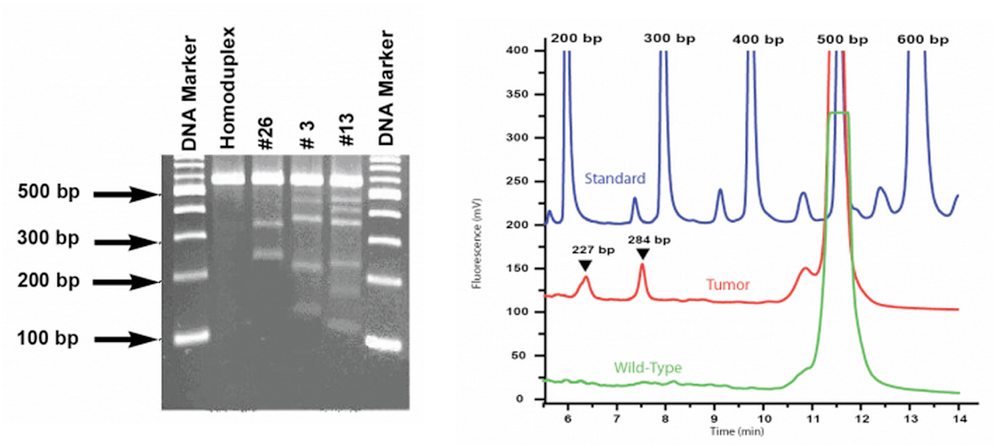
What are the advantages of enzymatic based methods of mutation scanning?
- Rapid and cheap method for screening for mutations
- Can analyse long range PCR products difficult to sequence
- Possible to detect multiple variants in single PCR products
- Automation feasible – might be useful for labs without high throughput Sequencing facilities.



